
- Homepage
- About the Joint Centre
- Research Areas & Projects
- Researchers
- News & Events
- Open Positions



A long standing problem of medical image analysis is the lack of annotated data (e.g. images of a tumour with its associated, expert annotated regions). Such data is crucially important for training automatic classification techniques which are able to automatically detect and delineate an anomaly in a new, previously unseen patient. Applications range from diagnosis, to more accurate treatment, from quantification of the effect of a drug to prevention.
Recent trends in machine learning and computer vision demonstrate that under certain constraints it is possible to generate annotated training data synthetically. The trained classifiers then are shown to work well on previously unseen test images. For instance, this technique is at the basis of the successful Microsoft Kinect sensor; where a body part classifier was trained on millions of synthetically generated images of people in various poses.
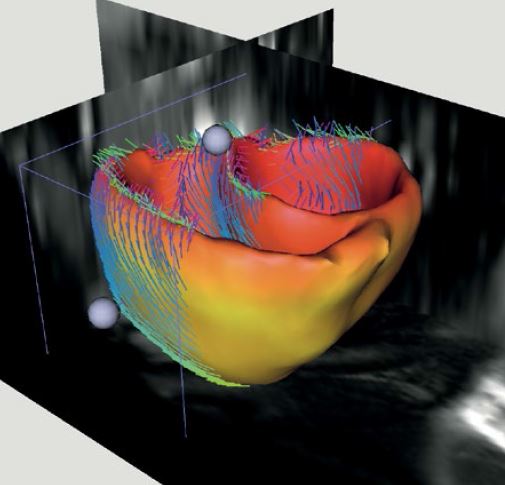
In this project we wish to do the same for medical images. This is very timely for a number of reasons. For instance, we now have available realistic generative models of e.g. brain scans, brain tumors, beating hearts, as well as full-body CT scans. So now the question is how well does a system trained on such synthetic images work on real patients scans, and how to modify the algorithm and its features to make it work more accurately.
As part of this project, the generated synthetic training and testing sets and their annotations will also be made available to the whole scientific community. This will enable various research teams to take part in competitions and compare their different algorithms on an equal base.
Neuroimaging is accumulating large functional MRI datasets that display, in spite of their low signal-to-noise ratio (SNR), brain activation patterns giving access to a meaningful representation of brain spatial organization. This ongoing accumulation is intensified via new large-scale international initiatives such as the Human Connectome Project or the recently accepted Human Brain Project, but also to existing open repositories of functional neuroimaging datasets. These datasets represent a very significant resource for the community, but require new analytic approaches in order to be fully exploited.
In order to provide a synthetic picture of the brain substrate of human cognition and its pathologies, we proceed by learning from large-scale datasets a brain atlas that summarizes adequately these functional activation maps drawing from a large number of protocols and subjects. Once learned, such an atlas is extremely useful to understand the large-scale functional organization of the brain: it is a tool for understanding brain segregation, the different encoding of many cognitive parameters into different brain regions, as well as brain integration, i.e. how remote brain regions co-activate across subjects and experiments.